AUCTORES
Globalize your Research
Review Article | DOI: https://doi.org/10.31579/2768-2757/156
1Medical microbiology department /College of Health Sciences/ Hawler Medical University/ Kurdistan region /Erbil/Iraq.
2Midwifery department, Erbil technical medical institute, Erbil polytechnic university, Kurdistan region/Erbil/ Iraq.
3Catholic university in Erbil.
*Corresponding Author: Fattma A. Ali., Medical microbiology department /College of Health Sciences/ Hawler Medical University/ Kurdistan region /Erbil/Iraq.
Citation: Fattma A. Ali, Khudhair Al-Daoody AA, Muna M. Najeeb, Media A. Othman, Maryam N. Philip, (2025), The Influence of Restriction Endonucleases for Cloning, Journal of Clinical Surgery and Research, 6(1); DOI:10.31579/2768-2757/156
Copyright: © 2025, Fattma A. Ali. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received: 18 November 2024 | Accepted: 16 December 2024 | Published: 14 January 2025
Keywords: restriction endonucleases; cloning; dna mapping; molecular biology; engineering
Background: Restriction//endonucleases, also referred to as restriction//enzymes, are enzymes that identify and cleave DNA at or in close proximity to certain DNA//sequences known as recognition//sites. Enzymes play a crucial role in molecular//biology by facilitating the manipulation of DNA, including tasks like cloning, gene//mapping, and DNA//sequencing. ). There are four primary categories of restriction//enzymes, namely types I,//II,//III//and//IV. These categories vary in terms of enzyme composition, //cofactor//needs, and mode//of//action.
Objectives: Our study aimed to carry out the role of the restriction endonucleases in the molecular microbiology mainly in cloning.
Main body: Historically, restriction//endonucleases were obtained in a pure form from the organism they naturally occur in. The advancement of gene//cloning//vectors and selection procedures facilitated the cloning//of //REases. Cloning facilitated the creation of huge amounts of meticulously purified enzymes and enabled the manipulation of restriction//endonucleases Presently. Traditional//cloning. When used together with DNA//ligases, REases///enable a reliable process of cutting and pasting DNA, allowing for the transfer of a specific DNA//fragment from one organism to another. Stanley Cohen and his colleagues utilized this technology to introduce foreign//DNA into native plasmids, resulting in the creation of cloning-plasmid vectors that can self-replicate in E.coli.
Conclusion: restriction endonucleases have greatly transformed the field of molecular biology by enabling progress in recombinant//DNA//technology, gene cloning, DNA//fingerprinting, and gene mapping. An example frequently encountered in molecular biology is the utilization of restriction//enzymes in the plasmid employed for the manufacture of human insulin. Restriction enzymes in plasmid DNA manipulation work by identifying and cutting certain DNA//sequences, called restriction sites, in the plasmid//DNA. Plasmids are compact, ring-shaped//DNA molecules that are frequently employed as carriers in molecular biology for the transportation and duplication of foreign DNA pieces. Restriction enzymes identify and attach to particular palindromic DNA sequences, referred to as restriction sites, inside the plasmid// DNA. Upon binding, these enzymes hydrolyze the plasmid //DNA at or in close proximity to their specific recognition sites, resulting in the production of linear DNA//fragments with either cohesive or blunt ends.
Restriction endonucleases, also referred to as restriction enzymes, are enzymes that identify and cleave DNA at or in close proximity to certain DNA sequences known as recognition sites. Enzymes play a crucial role in molecular biology by facilitating the manipulation of DNA, including tasks like cloning, gene mapping, and DNA sequencing [1] . Scientists faced difficulties in isolating and studying individual genes until the 1970s. The initial breakthrough involved the identification of restriction enzymes and the DNA ligase enzyme. The identification of this crucial breakthrough, together with the elucidation of scientific methodologies, empowered scientists to utilize these techniques for the purpose of isolating specific genes from a genome. An important breakthrough in the discipline occurred with the creation of plasmid cloning vectors, which enabled the reception and replication of isolated segments of DNA. The creation of these instruments resulted in the release of the initial recombinant DNA molecules in 1972.Genes are replicated by cloning techniques in order to facilitate the production of the proteins they carry, allowing for their abundant expression and subsequent purification. This is done for several purposes, including medical, experimental, and commercial uses [2]. Cloning in molecular biology is the act of creating several duplicates of a certain DNA sequence through the use of recombinant DNA technologies. The process entails the incorporation of a DNA segment of interest into a vector, such as a plasmid or a viral genome, and then amplifying it within a host cell, usually bacteria or yeast. Cloning allows for the separation, examination, and alteration of particular genes, which aids in a range of molecular biology tasks such as gene mapping, sequencing and functional analysis [3]. The elucidation of the mechanism by which restriction endonucleases, a group of bacterial enzymes, function, marked a significant advancement in the science of genetic engineering. Within a living organism, these enzymes participate in the recognition and fragmentation of foreign DNA that enters the cell. Therefore, their primary function is to safeguard the bacterium from phage infection. The relevant aspect for us is that these enzymes have the ability to identify specific DNA sequences. The enzymes utilized in DNA manipulation are specifically referred to as Class II restriction endonucleases. These enzymes cleave the DNA at a specific location within the recognition sequence. When DNA is treated with these enzymes, each molecule will be cut at the identical locations, resulting in the production of consistent pieces [4].Restriction endonucleases are a crucial part of restriction-modification (RM) systems, which primarily serve to safeguard bacterial cells against the intrusion of foreign DNA molecules [5]. There are 4 primary categories of restriction enzymes, namely types I, II, III and IV, as illustrated in table (1). These categories vary in terms of enzyme composition, cofactor needs, and mode of action [6].
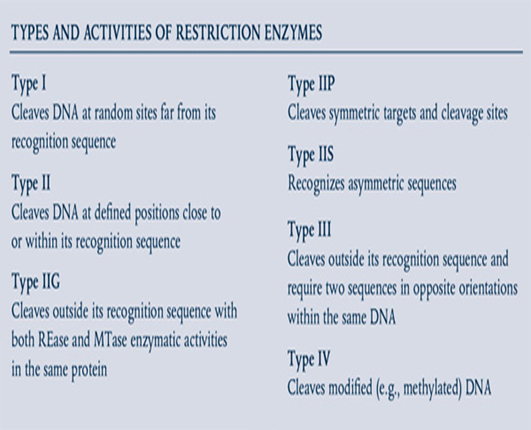

Table 1: types and activities of restriction enzymes.
The initial restriction endonucleases (REases) that were identified had the ability to recognize particular DNA sequences, but their cutting sites were varying distances away from the recognition sequence (referred to as Type I). Consequently, these enzymes were not very useful for DNA editing purposes. Following this, the identification and refinement of restriction endonucleases that could recognize and cleave specific DNA sequences (known as Type II REases) enabled scientists to carry out accurate modifications of DNA under laboratory conditions. This included tasks like inserting foreign genes into DNA and developing effective cloning tools. Currently, there are over 4,000 Restriction Endonucleases that have been identified. These enzymes are capable of detecting and binding to more than 300 different DNA sequences. The introduction of Polymerase Chain Reaction , real time-PCR, and PCR-based mutagenesis techniques revolutionized biological research in the subsequent decades. The type II representatives are the most extensively studied. They have a special ability to identify DNA sequences that are 4-8 base pairs long and can break the DNA either inside these sequences or in close proximity to them. Type II enzymes are extremely precise, making them essential for manipulating DNA [7]. Type II restriction endonucleases cleave specifically at the sites within their recognition sequences. Ligations of two strands cleaved by the same Type II enzyme are typically vulnerable to recognition and cleavage by the same enzyme. Therefore, it is necessary to remove that enzyme (via gel filtration or heat denaturation) before ligation takes place [8]. It is worth mentioning that there have been approximately 5000 restriction endonucleases identified so far, with nearly 4900 of them being type II enzymes [9] . Simultaneously, a total of 625 commercially accessible type II enzymes are capable of identifying only 239 distinct particular locations within DNA. Due to the limited number of specific sites available, continuous efforts are made to extensively search for new variations of restriction enzymes in natural sources and to artificially modify the specificity of known enzymes. While there are already hundreds of known DNA restriction enzymes , there is a need for enzymes that can recognize new DNA targets. Discovering new restriction enzymes that are highly specific for a unique nucleotide sequence at a "restriction site" and do not have nonspecific effects will increase the range of enzymes available for biotechnology. These new enzymes may also be able to replace existing ones and be more effective in their function [10].
2.1. Biotechnology of restriction enzymes (REases)
Historically, restriction endonucleases (REases) were obtained in a pure form from the organism they naturally occur in. The advancement of gene cloning vectors and selection procedures facilitated the cloning of REases. Cloning facilitated the creation of huge amounts of meticulously purified enzymes and enabled the manipulation of restriction endonucleases Presently [10].
2.2. Enhancing Performance in Engineering
Some restriction endonucleases have been discovered to exhibit cleavage activity at sites that are not their intended target sites, which is referred to as "star activity". This phenomenon has been well-documented. Among these enzymes, certain ones demonstrate star activity when subjected to less-than-ideal reaction circumstances, whilst others have a limited range of enzyme units that may fully digest a specific amount of substrate without showing any star activity. Scientists at NEB conducted extensive research to develop restriction enzymes that demonstrate limited or no star activity when used over extended periods of time and at large concentrations of the enzyme [11].
2.3. Developing novel sequence specificities in engineering
Efforts to modify them sequence specificities of Type IIP restriction endonuclease has mostly ineffective, perhaps due to the structural integration of sequence specificity determinant with them active sites of these enzymes. MmeI is a Type IIG restriction endonuclease that possesses both methyltransferase and REase activities within the same polypeptide. It specifically detects the target sequence TCCRAC using the target recognition domain located within its methyltransferase component . This gave a great chance to manipulate the REase's sequence specificity. The sharing of target recognition domain between them REasen and methyltransferase activities has the additional benefit of causing an equal change in methyltransferase activity regardless of the specific target sequence cleavage ,This protects the new target site from being cleaved in recombinant host cells. By conducting bioinformatics analysis on protein sequences that are similar, researchers at NEB were able to identify the precise amino acid residues that can recognize particular bases in the target sequences. They then developed MmeI mutants that had modified specificities in terms of sequence recognition [12].
2.4. Engineering Nicking Endonucleases
Research focused on restriction endonucleases has uncovered unexpected discoveries regarding the apparently simple process of cleavage. Typically, Type IIP REases function as homo dimers, where each monomer cleaves one half of the palindromic site. On the contrary, Type IIS restriction endonucleases demonstrate a wide variety of ways for cutting both strands of DNA. These mechanisms include hetero dimerization, as seen in enzymes like BtsI and BbvCI as well as sequential cleavage of the double-stranded DNA as a single molecule, as seen in the case of FokI. The unique characteristics of these traits have been utilized to develop strand.-specific nicking enzymes (NEases) [13].
2.5. Applications involving the use of restriction enzymes include:
2.5.1. Traditional cloning.
When used together with DNA ligases, REases enable a reliable process of cutting and pasting DNA, allowing for the transfer of a specific DNA fragment from one organism to another (Figure 1). Stanley Cohen and his colleagues utilized this technology to introduce foreign DNA into native plasmids, resulting in the creation of cloning.-plasmid vectors that can self-replicate in E.coli. These vectors have become essential tools in modern biology, allowing for the cloning of DNA and the synthesis of recombinant proteins. Restriction enzymes serve as valuable instruments for post-cloning confirmation, ensuring the accurate occurrence of insertions. The conventional process of cloning, in conjunction with DNA amplification techniques like PCR and RT-PCR, has become a widely used method for studying molecular mechanisms, particularly in relation to restriction endonucleases [14].
Figure 1. Traditional Cloning Workflow
PCR is employed to introduce restriction sites at both ends of a double-stranded DNA molecule. Subsequently, the DNA molecule is cleaved by the matching restriction enzymes. The fragmented DNA can subsequently be joined to a plasmid vector that has been fragmented by the same or compatible restriction endonucleases using T4 DNA ligase. DNA fragments can be transferred across vectors by digesting them using restriction endonucleases and then ligating them to the compatible ends of the target vector.
2.5.2. DNA Mapping
Daniel Nathans, armed with a limited number of restriction endonucleases in the earlym1970s, successfully identified the functional units of SV40 DNA. This marked the beginning of the era of "restriction mapping" and the comparison of complicated genomes. Over time, it has developed into advanced techniques that enable the identification of individual variations in DNA sequences and the presence of genetic mutations. These methods are used for various purposes such as identifying locations associated with genetic disorders, evaluating the genetic variability within populations, and conducting parental testing [15].
2.5.3. Understanding Epigenetic Modifications
The heightened responsiveness of REases to the methylation state of certain bases has been utilized to accurately locate changed bases in genomes. Restriction Landmark Genome Scanning is a mapping approach that uses 2-dimensional gel electrophoresis and specific enzymes (NotI, AscI, EagI, or BssHII) to analyze alterations in the methylation patterns of the genome in normal and cancer cells. Methylation.-Sensitive Amplification is a technique used to amplify DNA fragments that are sensitive to methylation. Polymorphism is utilized by using the varying sensitivity of MspI and HpaII enzymes towards the methylation status of the second C in the sequence CCGG. This allows for the identification of 5-methylcytosine or 5-hydroxymethylcytosine[16]. Furthermore, the recently discovered restriction enzymes (REases) MspJI, FspEI, and LpnPI have the ability to recognize and cleave DNA at specific sites containing 5-methylcytosine and 5-hydroxymethylcytosine. Additionally, there are enzymes such as PvuRts1I and AbaSI that have a preference for cleaving 5-hydroxymethylcytosine or 5-ghmC rather than 5-methylcytosine or C. These enzymes [17], have the potential to be used for efficiently mapping cytosine-based epigenetic markers in genomes that have undergone cytosine methylation [18].
2.5.4. In vitro DNA Assembly Technologies
Synthetic biology is an expanding discipline that utilizes specific elements to construct biological systems for the purpose of investigating biological processes and developing practical biological devices. Novel technologies like BioBrick™ were first developed to simplify the construction of biological systems. In recent times, the synthetic biology community has extensively embraced more resilient methods, such as Golden Gate. Assembly and Gibson Assembly™. Both methods enable the simultaneous and smooth joining of numerous DNA fragments without the need for unconventional bases [19]. The BioBricks group aimed to develop a large collection of "standardized parts" of DNA molecules to facilitate quick gene assembly. The biennial International Genetically Engineered Machines competition fostered the growth of the BioBricks community and generated significant interest among university students in the field of synthetic biology. Utilizing the conventional REase-ligation technique. However, they do result in the presence of scar sequences at the junctions. In order to develop a functional biological system, it is necessary to undergo numerous cloning cycles [20].
2.5.5. Golden Gate Assembly
Golden Gate Assembly and its derivative technologies utilize the capacity of Type IIS restriction endonucleases to cleave DNA outside of the recognition sequence . The inserts and cloning vectors are specifically designed to position the Type IIS recognition site away from the cleavage site, allowing the Type IIS REase to eliminate the recognition sequence from the assembly (Figure 2). There are three advantages to such an arrangement:
1. The sequence that extends beyond the restriction enzyme recognition site is not determined by the restriction enzyme, and as a result, no more sequence is added.
2.The precise sequencing of the overhangs enables the systematic assembly of numerous fragments at the same time.
3. The ligated product undergoes elimination of the restriction site, allowing for simultaneous digestion and ligation [21].
The ultimate outcome is the organized and uninterrupted combination of DNA fragments in a single reaction. The precision of the assembly relies on the length of the overhang sequences .Hence, Type IIS restriction endonucleases that generate four-bases overhangs (such as BsaI/BsaI-HF, BbsI, BsmBI, and Esp3I) are favored. A limitation of these Type IIS REase-based approaches is that the presence of a limited number of overhanging bases can result in the incorrect joining of fragments that have comparable overhang sequences [22] . Additionally, it is crucial to confirm that Type IIS REase sites utilized are absent in fragments used for assembling the intended result. However, Golden Gate Assembly is a reliable and efficient approach that produces several targeted alterations [23] and combines various DNA fragments [24] . With the growing availability of open source methodologies and reagents, the use of Golden Gate Assembly has become widespread in the creation of custom-specific TALENs for in vivo gene editing, among other applications [25].
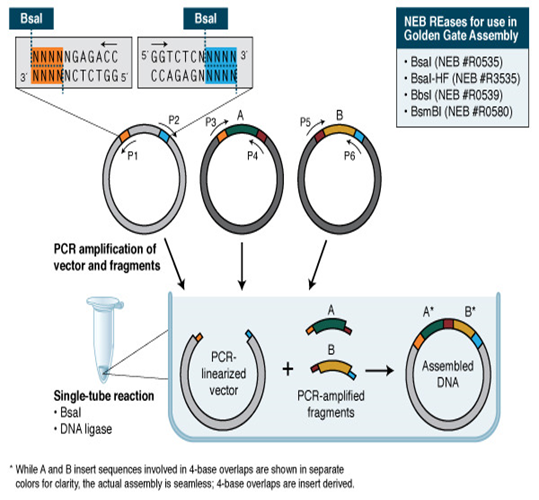
Figure 2: Golden Gate Assembly Workflow.
Golden Gate Assembly involves the addition of a BsaI recognition site (GGTCTC) to both ends of a double-stranded DNA fragment located away from the cleavage site. This allows for the removal of the BsaI site by digesting it with BsaI or BsaI-HF (GGTCTC 1/5). After being split, the protruding sequences of the adjacent fragments bind together. DNA ligase then closes the gaps in order to generate a novel DNA molecule that is chemically bonded. It is possible to concurrently cut and ligate multiple sections of DNA.
2.5.6. Gibson Assembly
Daniel G. Gibson, from the J. Craig Venter Institute, explained a reliable technique that uses exonucleases to assemble DNA without any gaps and in the proper sequence. The reaction is performed at a constant temperature utilizing three enzymatic processes: a 5' exonuclease creates extended ends, a polymerase fills in the gaps of the single-stranded areas that have been joined together, and a DNA ligase closes the gaps that have been joined and filled in (Figure 3). The process involved assembling the mouse mitochondrial genome, which has a size of 16.3 kb, using 600 overlapping 60-mers [26]. Gibson Assembly was employed alongside in vivo assembly in yeast to synthesize the 1.1 Mbp Mycoplasma mycoides genome. A recipient cell of M. capricolum was created by transplanting the synthesized genome, resulting in the production of new self-replicating M. mycoides cells. Gibson Assembly can also be employed for cloning purposes. This involves combining a DNA insert with a vector that has been digested by restriction enzymes, followed by a process called transformation. Using the Gibson Assembly Cloning Kit, this entire procedure can be accomplished in just under two hours. Gibson Assembly has various other uses, such as facilitating the incorporation of numerous mutations, constructing plasmid vectors using chemically genera oligonucleotides, and generating libraries of genes and pathways with combinatorial diversity [27].
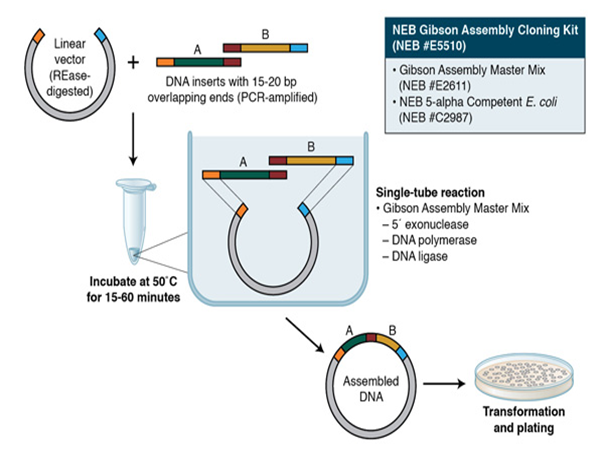
Figure 3: Gibson Assembly Workflow.
Gibson Assembly utilizes a combination of three enzymatic activities in a single-tube reaction: 5´mexonuclease, the 3´ extension activity of a DNA polymerase, and DNA ligase activity. The 5´exonucleasemactivity removes nucleotides off the 5´ end of the sequence, revealing the complementary sequence for annealing. The polymerase activity subsequently completes the missing sections on the areas that have been annealed. Following the nick, a DNA ligase enzymatically plugs the gap and forms covalent bonds between the DNA fragments, effectively joining them together. The length of the overlapping sequence of adjacent fragments is significantly greater than that employed in Golden Gate Assembly, leading to a higher proportion of accurate assemblies. The NEB Gibson Assembly Master Mix and Gibson Assembly Cloning Kit facilitate fast assembly at a temperature of 50˚C.
2.5.7. Construction of DNA Libraries
The technique of Serial Analysis of Gene Expression has facilitated the detection and measurement of a substantial quantity of RNA transcripts. It has been extensively utilized in cancer research to detect mutations and investigate gene expression. Restriction enzymes play a crucial role in the Serial Analysis of Gene Expression procedure. NlaIII serves as a crucial anchoring enzyme due to its distinctive ability to identify a four-bp sequence CATG and generate a four nucleotide overhang with the same sequence. Type IIS enzymes, such as FokI, BsmFI, MmeI, and EcoP15I, are used as tagging enzymes in Serial Analysis of Gene Expression analyses. These enzymes cleave DNA sequences that are located further away from the recognition sequence. This allows for a higher amount of information to be obtained in SAGE analyses. Examples of SAGE studies that have utilized Type IIS enzymes include [28] for FokI and BsmFI [29] for MmeI [30] for EcoP15I for Deep Serial Analysis of Gene Expression [31] .
Chromosome conformation capture (3C) and related techniques enable the precise mapping of the spatial arrangements of genomes with exceptional resolution and efficiency. REases are essential for generating compatible DNA ends that are cross-linked to their interacting proteins.
This allows for the ligation and identification of spatially related sequences by high-throughput sequencing. Despite the fact that restriction endonucleases cannot facilitate the random fragmentation of DNA, they are being utilized in innovative target enrichment techniques such as hairpin adaptor ligation and HaloPlex™ enrichment (Agilent) [32]. The long-reach REase, AcuI, and USER™ Enzyme are utilized for the purpose of incorporating tags into sample DNA. This DNA is subsequently amplified through rolling circle amplification, resulting in the formation of long, single-stranded DNA "nanoballs". These nanoballs serve as templates in the high density, chip-based sequencing-by-ligation methodology, which was developed by Complete Genomics [33]. The DNA library for genotyping-by-sequencing technique in the investigation of maize sequence diversity was generated using ApeKI [34].
2.5.8. Creation of Nicks in DNA
Prior to the availability of NEases, non-hydrolyzable phosphorothioate groups were added to a particular strand of the target DNA. This allowed REases to create specific nicks in the DNA, based on the sequence and strand. These nicks were used in applications like strand displacement amplification, where a DNA polymerase (such as Bstm2.0 DNA Polymerase) extends from the newly formed 3'-hydroxyl end and replicates the complementary sequence [35]. Repeated rounds of nicking and extension amplify specific single-stranded portions of the sample DNA without requiring thermocycling, as the nicking site is regenerated. Nucleases significantly simplify the process of such applications and enable the development of applications that cannot be accomplished by restriction endonucleases. The utilization of nicking enzyme-based isothermal DNA amplification technologies, such as RCA, NESA, EXPAR and other comparable amplification systems, has demonstrated the ability to detect extremely low quantities of DNA [36]. The technique of nicking-based DNA amplification has been integrated in to molecular beacon technologies in order to enhance the signal amplification [37]. Implementing these sample and or signal amplification approaches can result in straightforward yet very sensitive and specific methods for detecting target DNA molecules in the field (NEAR, EnviroLogix™). The process involves attaching adaptors with nicking sites to the ends of blunt-ended DNA. This allows for the rapid amplification of a specific region of double-stranded DNA by the combined actions of NEases and strand-displacing DNA polymerase [38] .Nicking-extension cycling amplification is capable of multiplexing and has the potential to attain greater fidelity than PCR. The collective action of NEases and Bst DNA polymerase has been utilized to include specific fluorescent markers into long chromosomal DNA in vitro for the purpose of viewing (nanocoding) [39] . The utilization of nicking enzymes in novel ways includes the production of reporter plasmids with altered bases or structures and the development of a DNA motor that delivers DNA payload without the need for additional energy [40].
2.5.9. In vivo Gene Editing
The capacity to manipulate DNA using restriction endonucleases in a laboratory setting has naturally prompted efforts to achieve the same process within living organisms, with the aim of rectifying genetic mutations responsible for causing hereditary disorders. The utilization of restriction endonucleases and homing endonucleases in Restriction Enzyme Mediated Integration (REMI) enabled the creation of transgenic embryos in higher organisms [41]. Unfortunately, there is no regulation or supervision of the integration site. Zinc Finger Nucleases and Transcription Activator-like Effector Nucleases have been successfully used to edit genes by creating precise double stranded breaks in complicated genomes. The success of gene editing in model organisms and livestock has led to the testing of its therapeutic potential in Phase I/II clinical trials. These trials involve the use of Zinc Finger Nucleases to improve T helper cells counts by suppressing the expression of the CCR5 gene in autologous T-cells from HIV patients. The clinical trial with this approach has been registered under the identifier NCT00842634 on ClinicalTrials.gov. The latest studies on CRISPR, the adaptive defense mechanism found in bacteria and archaea, have shown the capabilities of the Cas9-crRNA complex as programmable RNA-guided DNA endonucleases and strand-specific nicking endonucleases for gene editing within living organisms [42].
2.6. Moving forward
Restriction enzymes have played a significant role in facilitating gene cloning and revolutionizing the field of molecular biology. Emerging technologies like Golden Gate Assembly and Gibson Assembly are continuously advancing our capacity to generate novel DNA molecules. The ability to establish new recognition specificity in the/ MmeI family restriction endonucleases , the development of additional non-specific endonucleases and the identification of more modification-specific REases, are continuously leading to the creation of novel tools for DNA manipulation and study of the epigenome. The utilization of these enzymes in novel ways will expand the role of REases beyond molecular cloning, further expediting the advancement of biotechnology and introducing us to fresh prospects and obstacles [10].
2.7. The commercialization and availability of restriction enzymes
The commercialization and accessibility of restriction enzymes have had a substantial impact on the progress of molecular biology research. Due to the growing need for these enzymes in different fields, numerous businesses have successfully brought a diverse array of restriction enzymes to the market, offering researchers easy access to top-notch reagents. The commercialization of restriction enzymes commenced during the initial stages of molecular biology research, as corporations identified the need for these indispensable instruments. Currently, there is a diverse selection of restriction enzymes that can be purchased from several providers, including New England Biolabs, Thermo Fisher Scientific, and Promega Corporation. The companies New England Biolabs, Thermo Fisher Scientific, and Promega Corporation provide a diverse range of restriction enzymes. These enzymes have distinct recognition sequences, cleavage patterns, and optimal reaction conditions. As a result, researchers have access to a comprehensive set of tools for manipulating and analyzing DNA [1].
In conclusion restriction endonucleases, also known as restriction enzymes, are fundamental instruments in the field of molecular biology, serving as essential agents in the manipulation, analysis and cloning of DNA. These enzymes have the ability to identify and cut DNA at particular nucleotide sequences, allowing for accurate manipulation of DNA fragmentation and recombination. Since being discovered in the late 1960s, restriction endonucleases have greatly transformed the field of molecular biology by enabling progress in recombinant DNA technology, gene cloning, DNA fingerprinting, and gene mapping. An example frequently encountered in molecular biology is the utilization of restriction enzymes in the plasmid employed for the manufacture of human insulin. Restriction enzymes in plasmid DNA manipulation work by identifying and cutting certain DNA sequences, called restriction sites, in the plasmid DNA. Plasmids are compact, ring-shaped DNA molecules that are frequently employed as carriers in molecular biology for the transportation and duplication of foreign DNA pieces. Restriction enzymes identify and attach to particular palindromic DNA sequences, referred to as restriction sites, inside the plasmid DNA. Upon binding, these enzymes hydrolyze the plasmid DNA at or in close proximity to their specific recognition sites, resulting in the production of linear DNA fragments with either cohesive or blunt ends. Subsequently, the linearized plasmid DNA can be merged with a DNA fragment of interest, and then joined together using DNA ligase, resulting in the formation of a recombinant plasmid. The recombinant plasmid can be placed into a host organism, such as bacteria, where it can duplicate and produce the inserted DNA fragment. Restriction enzymes possess a high degree of specificity and precision when it comes to cutting DNA sequences. This characteristic allows for accurate manipulation and cloning of DNA fragments within plasmid vectors. As a result, different applications in molecular biology, such as gene cloning, gene expression research, and functional genomics, are made possible. The broad use and importance of restriction enzymes in research laboratories worldwide is further emphasized by the fact that they are commercially available from a variety of sources. Restriction endonucleases are essential tools in molecular biology that play a crucial role in driving innovation and making discoveries in biological research. They contribute significantly to our understanding of genetics, disease mechanisms, and the creation of new therapeutic approaches.
Clearly Auctoresonline and particularly Psychology and Mental Health Care Journal is dedicated to improving health care services for individuals and populations. The editorial boards' ability to efficiently recognize and share the global importance of health literacy with a variety of stakeholders. Auctoresonline publishing platform can be used to facilitate of optimal client-based services and should be added to health care professionals' repertoire of evidence-based health care resources.

Journal of Clinical Cardiology and Cardiovascular Intervention The submission and review process was adequate. However I think that the publication total value should have been enlightened in early fases. Thank you for all.

Journal of Women Health Care and Issues By the present mail, I want to say thank to you and tour colleagues for facilitating my published article. Specially thank you for the peer review process, support from the editorial office. I appreciate positively the quality of your journal.
Journal of Clinical Research and Reports I would be very delighted to submit my testimonial regarding the reviewer board and the editorial office. The reviewer board were accurate and helpful regarding any modifications for my manuscript. And the editorial office were very helpful and supportive in contacting and monitoring with any update and offering help. It was my pleasure to contribute with your promising Journal and I am looking forward for more collaboration.

We would like to thank the Journal of Thoracic Disease and Cardiothoracic Surgery because of the services they provided us for our articles. The peer-review process was done in a very excellent time manner, and the opinions of the reviewers helped us to improve our manuscript further. The editorial office had an outstanding correspondence with us and guided us in many ways. During a hard time of the pandemic that is affecting every one of us tremendously, the editorial office helped us make everything easier for publishing scientific work. Hope for a more scientific relationship with your Journal.

The peer-review process which consisted high quality queries on the paper. I did answer six reviewers’ questions and comments before the paper was accepted. The support from the editorial office is excellent.

Journal of Neuroscience and Neurological Surgery. I had the experience of publishing a research article recently. The whole process was simple from submission to publication. The reviewers made specific and valuable recommendations and corrections that improved the quality of my publication. I strongly recommend this Journal.

Dr. Katarzyna Byczkowska My testimonial covering: "The peer review process is quick and effective. The support from the editorial office is very professional and friendly. Quality of the Clinical Cardiology and Cardiovascular Interventions is scientific and publishes ground-breaking research on cardiology that is useful for other professionals in the field.

Thank you most sincerely, with regard to the support you have given in relation to the reviewing process and the processing of my article entitled "Large Cell Neuroendocrine Carcinoma of The Prostate Gland: A Review and Update" for publication in your esteemed Journal, Journal of Cancer Research and Cellular Therapeutics". The editorial team has been very supportive.

Testimony of Journal of Clinical Otorhinolaryngology: work with your Reviews has been a educational and constructive experience. The editorial office were very helpful and supportive. It was a pleasure to contribute to your Journal.

Dr. Bernard Terkimbi Utoo, I am happy to publish my scientific work in Journal of Women Health Care and Issues (JWHCI). The manuscript submission was seamless and peer review process was top notch. I was amazed that 4 reviewers worked on the manuscript which made it a highly technical, standard and excellent quality paper. I appreciate the format and consideration for the APC as well as the speed of publication. It is my pleasure to continue with this scientific relationship with the esteem JWHCI.

This is an acknowledgment for peer reviewers, editorial board of Journal of Clinical Research and Reports. They show a lot of consideration for us as publishers for our research article “Evaluation of the different factors associated with side effects of COVID-19 vaccination on medical students, Mutah university, Al-Karak, Jordan”, in a very professional and easy way. This journal is one of outstanding medical journal.
Dear Hao Jiang, to Journal of Nutrition and Food Processing We greatly appreciate the efficient, professional and rapid processing of our paper by your team. If there is anything else we should do, please do not hesitate to let us know. On behalf of my co-authors, we would like to express our great appreciation to editor and reviewers.

As an author who has recently published in the journal "Brain and Neurological Disorders". I am delighted to provide a testimonial on the peer review process, editorial office support, and the overall quality of the journal. The peer review process at Brain and Neurological Disorders is rigorous and meticulous, ensuring that only high-quality, evidence-based research is published. The reviewers are experts in their fields, and their comments and suggestions were constructive and helped improve the quality of my manuscript. The review process was timely and efficient, with clear communication from the editorial office at each stage. The support from the editorial office was exceptional throughout the entire process. The editorial staff was responsive, professional, and always willing to help. They provided valuable guidance on formatting, structure, and ethical considerations, making the submission process seamless. Moreover, they kept me informed about the status of my manuscript and provided timely updates, which made the process less stressful. The journal Brain and Neurological Disorders is of the highest quality, with a strong focus on publishing cutting-edge research in the field of neurology. The articles published in this journal are well-researched, rigorously peer-reviewed, and written by experts in the field. The journal maintains high standards, ensuring that readers are provided with the most up-to-date and reliable information on brain and neurological disorders. In conclusion, I had a wonderful experience publishing in Brain and Neurological Disorders. The peer review process was thorough, the editorial office provided exceptional support, and the journal's quality is second to none. I would highly recommend this journal to any researcher working in the field of neurology and brain disorders.

Dear Agrippa Hilda, Journal of Neuroscience and Neurological Surgery, Editorial Coordinator, I trust this message finds you well. I want to extend my appreciation for considering my article for publication in your esteemed journal. I am pleased to provide a testimonial regarding the peer review process and the support received from your editorial office. The peer review process for my paper was carried out in a highly professional and thorough manner. The feedback and comments provided by the authors were constructive and very useful in improving the quality of the manuscript. This rigorous assessment process undoubtedly contributes to the high standards maintained by your journal.

International Journal of Clinical Case Reports and Reviews. I strongly recommend to consider submitting your work to this high-quality journal. The support and availability of the Editorial staff is outstanding and the review process was both efficient and rigorous.

Thank you very much for publishing my Research Article titled “Comparing Treatment Outcome Of Allergic Rhinitis Patients After Using Fluticasone Nasal Spray And Nasal Douching" in the Journal of Clinical Otorhinolaryngology. As Medical Professionals we are immensely benefited from study of various informative Articles and Papers published in this high quality Journal. I look forward to enriching my knowledge by regular study of the Journal and contribute my future work in the field of ENT through the Journal for use by the medical fraternity. The support from the Editorial office was excellent and very prompt. I also welcome the comments received from the readers of my Research Article.

Dear Erica Kelsey, Editorial Coordinator of Cancer Research and Cellular Therapeutics Our team is very satisfied with the processing of our paper by your journal. That was fast, efficient, rigorous, but without unnecessary complications. We appreciated the very short time between the submission of the paper and its publication on line on your site.

I am very glad to say that the peer review process is very successful and fast and support from the Editorial Office. Therefore, I would like to continue our scientific relationship for a long time. And I especially thank you for your kindly attention towards my article. Have a good day!

"We recently published an article entitled “Influence of beta-Cyclodextrins upon the Degradation of Carbofuran Derivatives under Alkaline Conditions" in the Journal of “Pesticides and Biofertilizers” to show that the cyclodextrins protect the carbamates increasing their half-life time in the presence of basic conditions This will be very helpful to understand carbofuran behaviour in the analytical, agro-environmental and food areas. We greatly appreciated the interaction with the editor and the editorial team; we were particularly well accompanied during the course of the revision process, since all various steps towards publication were short and without delay".

I would like to express my gratitude towards you process of article review and submission. I found this to be very fair and expedient. Your follow up has been excellent. I have many publications in national and international journal and your process has been one of the best so far. Keep up the great work.

We are grateful for this opportunity to provide a glowing recommendation to the Journal of Psychiatry and Psychotherapy. We found that the editorial team were very supportive, helpful, kept us abreast of timelines and over all very professional in nature. The peer review process was rigorous, efficient and constructive that really enhanced our article submission. The experience with this journal remains one of our best ever and we look forward to providing future submissions in the near future.

I am very pleased to serve as EBM of the journal, I hope many years of my experience in stem cells can help the journal from one way or another. As we know, stem cells hold great potential for regenerative medicine, which are mostly used to promote the repair response of diseased, dysfunctional or injured tissue using stem cells or their derivatives. I think Stem Cell Research and Therapeutics International is a great platform to publish and share the understanding towards the biology and translational or clinical application of stem cells.

I would like to give my testimony in the support I have got by the peer review process and to support the editorial office where they were of asset to support young author like me to be encouraged to publish their work in your respected journal and globalize and share knowledge across the globe. I really give my great gratitude to your journal and the peer review including the editorial office.

I am delighted to publish our manuscript entitled "A Perspective on Cocaine Induced Stroke - Its Mechanisms and Management" in the Journal of Neuroscience and Neurological Surgery. The peer review process, support from the editorial office, and quality of the journal are excellent. The manuscripts published are of high quality and of excellent scientific value. I recommend this journal very much to colleagues.

Dr.Tania Muñoz, My experience as researcher and author of a review article in The Journal Clinical Cardiology and Interventions has been very enriching and stimulating. The editorial team is excellent, performs its work with absolute responsibility and delivery. They are proactive, dynamic and receptive to all proposals. Supporting at all times the vast universe of authors who choose them as an option for publication. The team of review specialists, members of the editorial board, are brilliant professionals, with remarkable performance in medical research and scientific methodology. Together they form a frontline team that consolidates the JCCI as a magnificent option for the publication and review of high-level medical articles and broad collective interest. I am honored to be able to share my review article and open to receive all your comments.

“The peer review process of JPMHC is quick and effective. Authors are benefited by good and professional reviewers with huge experience in the field of psychology and mental health. The support from the editorial office is very professional. People to contact to are friendly and happy to help and assist any query authors might have. Quality of the Journal is scientific and publishes ground-breaking research on mental health that is useful for other professionals in the field”.

Dear editorial department: On behalf of our team, I hereby certify the reliability and superiority of the International Journal of Clinical Case Reports and Reviews in the peer review process, editorial support, and journal quality. Firstly, the peer review process of the International Journal of Clinical Case Reports and Reviews is rigorous, fair, transparent, fast, and of high quality. The editorial department invites experts from relevant fields as anonymous reviewers to review all submitted manuscripts. These experts have rich academic backgrounds and experience, and can accurately evaluate the academic quality, originality, and suitability of manuscripts. The editorial department is committed to ensuring the rigor of the peer review process, while also making every effort to ensure a fast review cycle to meet the needs of authors and the academic community. Secondly, the editorial team of the International Journal of Clinical Case Reports and Reviews is composed of a group of senior scholars and professionals with rich experience and professional knowledge in related fields. The editorial department is committed to assisting authors in improving their manuscripts, ensuring their academic accuracy, clarity, and completeness. Editors actively collaborate with authors, providing useful suggestions and feedback to promote the improvement and development of the manuscript. We believe that the support of the editorial department is one of the key factors in ensuring the quality of the journal. Finally, the International Journal of Clinical Case Reports and Reviews is renowned for its high- quality articles and strict academic standards. The editorial department is committed to publishing innovative and academically valuable research results to promote the development and progress of related fields. The International Journal of Clinical Case Reports and Reviews is reasonably priced and ensures excellent service and quality ratio, allowing authors to obtain high-level academic publishing opportunities in an affordable manner. I hereby solemnly declare that the International Journal of Clinical Case Reports and Reviews has a high level of credibility and superiority in terms of peer review process, editorial support, reasonable fees, and journal quality. Sincerely, Rui Tao.

Clinical Cardiology and Cardiovascular Interventions I testity the covering of the peer review process, support from the editorial office, and quality of the journal.

Clinical Cardiology and Cardiovascular Interventions, we deeply appreciate the interest shown in our work and its publication. It has been a true pleasure to collaborate with you. The peer review process, as well as the support provided by the editorial office, have been exceptional, and the quality of the journal is very high, which was a determining factor in our decision to publish with you.
The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews journal clinically in the future time.

Clinical Cardiology and Cardiovascular Interventions, I would like to express my sincerest gratitude for the trust placed in our team for the publication in your journal. It has been a true pleasure to collaborate with you on this project. I am pleased to inform you that both the peer review process and the attention from the editorial coordination have been excellent. Your team has worked with dedication and professionalism to ensure that your publication meets the highest standards of quality. We are confident that this collaboration will result in mutual success, and we are eager to see the fruits of this shared effort.

Dear Dr. Jessica Magne, Editorial Coordinator 0f Clinical Cardiology and Cardiovascular Interventions, I hope this message finds you well. I want to express my utmost gratitude for your excellent work and for the dedication and speed in the publication process of my article titled "Navigating Innovation: Qualitative Insights on Using Technology for Health Education in Acute Coronary Syndrome Patients." I am very satisfied with the peer review process, the support from the editorial office, and the quality of the journal. I hope we can maintain our scientific relationship in the long term.
Dear Monica Gissare, - Editorial Coordinator of Nutrition and Food Processing. ¨My testimony with you is truly professional, with a positive response regarding the follow-up of the article and its review, you took into account my qualities and the importance of the topic¨.

Dear Dr. Jessica Magne, Editorial Coordinator 0f Clinical Cardiology and Cardiovascular Interventions, The review process for the article “The Handling of Anti-aggregants and Anticoagulants in the Oncologic Heart Patient Submitted to Surgery” was extremely rigorous and detailed. From the initial submission to the final acceptance, the editorial team at the “Journal of Clinical Cardiology and Cardiovascular Interventions” demonstrated a high level of professionalism and dedication. The reviewers provided constructive and detailed feedback, which was essential for improving the quality of our work. Communication was always clear and efficient, ensuring that all our questions were promptly addressed. The quality of the “Journal of Clinical Cardiology and Cardiovascular Interventions” is undeniable. It is a peer-reviewed, open-access publication dedicated exclusively to disseminating high-quality research in the field of clinical cardiology and cardiovascular interventions. The journal's impact factor is currently under evaluation, and it is indexed in reputable databases, which further reinforces its credibility and relevance in the scientific field. I highly recommend this journal to researchers looking for a reputable platform to publish their studies.
Dear Editorial Coordinator of the Journal of Nutrition and Food Processing! "I would like to thank the Journal of Nutrition and Food Processing for including and publishing my article. The peer review process was very quick, movement and precise. The Editorial Board has done an extremely conscientious job with much help, valuable comments and advices. I find the journal very valuable from a professional point of view, thank you very much for allowing me to be part of it and I would like to participate in the future!”

Dealing with The Journal of Neurology and Neurological Surgery was very smooth and comprehensive. The office staff took time to address my needs and the response from editors and the office was prompt and fair. I certainly hope to publish with this journal again.Their professionalism is apparent and more than satisfactory. Susan Weiner

My Testimonial Covering as fellowing: Lin-Show Chin. The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews.

My experience publishing in Psychology and Mental Health Care was exceptional. The peer review process was rigorous and constructive, with reviewers providing valuable insights that helped enhance the quality of our work. The editorial team was highly supportive and responsive, making the submission process smooth and efficient. The journal's commitment to high standards and academic rigor makes it a respected platform for quality research. I am grateful for the opportunity to publish in such a reputable journal.
My experience publishing in International Journal of Clinical Case Reports and Reviews was exceptional. I Come forth to Provide a Testimonial Covering the Peer Review Process and the editorial office for the Professional and Impartial Evaluation of the Manuscript.

I would like to offer my testimony in the support. I have received through the peer review process and support the editorial office where they are to support young authors like me, encourage them to publish their work in your esteemed journals, and globalize and share knowledge globally. I really appreciate your journal, peer review, and editorial office.
Dear Agrippa Hilda- Editorial Coordinator of Journal of Neuroscience and Neurological Surgery, "The peer review process was very quick and of high quality, which can also be seen in the articles in the journal. The collaboration with the editorial office was very good."

I would like to express my sincere gratitude for the support and efficiency provided by the editorial office throughout the publication process of my article, “Delayed Vulvar Metastases from Rectal Carcinoma: A Case Report.” I greatly appreciate the assistance and guidance I received from your team, which made the entire process smooth and efficient. The peer review process was thorough and constructive, contributing to the overall quality of the final article. I am very grateful for the high level of professionalism and commitment shown by the editorial staff, and I look forward to maintaining a long-term collaboration with the International Journal of Clinical Case Reports and Reviews.
To Dear Erin Aust, I would like to express my heartfelt appreciation for the opportunity to have my work published in this esteemed journal. The entire publication process was smooth and well-organized, and I am extremely satisfied with the final result. The Editorial Team demonstrated the utmost professionalism, providing prompt and insightful feedback throughout the review process. Their clear communication and constructive suggestions were invaluable in enhancing my manuscript, and their meticulous attention to detail and dedication to quality are truly commendable. Additionally, the support from the Editorial Office was exceptional. From the initial submission to the final publication, I was guided through every step of the process with great care and professionalism. The team's responsiveness and assistance made the entire experience both easy and stress-free. I am also deeply impressed by the quality and reputation of the journal. It is an honor to have my research featured in such a respected publication, and I am confident that it will make a meaningful contribution to the field.

"I am grateful for the opportunity of contributing to [International Journal of Clinical Case Reports and Reviews] and for the rigorous review process that enhances the quality of research published in your esteemed journal. I sincerely appreciate the time and effort of your team who have dedicatedly helped me in improvising changes and modifying my manuscript. The insightful comments and constructive feedback provided have been invaluable in refining and strengthening my work".

I thank the ‘Journal of Clinical Research and Reports’ for accepting this article for publication. This is a rigorously peer reviewed journal which is on all major global scientific data bases. I note the review process was prompt, thorough and professionally critical. It gave us an insight into a number of important scientific/statistical issues. The review prompted us to review the relevant literature again and look at the limitations of the study. The peer reviewers were open, clear in the instructions and the editorial team was very prompt in their communication. This journal certainly publishes quality research articles. I would recommend the journal for any future publications.

Dear Jessica Magne, with gratitude for the joint work. Fast process of receiving and processing the submitted scientific materials in “Clinical Cardiology and Cardiovascular Interventions”. High level of competence of the editors with clear and correct recommendations and ideas for enriching the article.

We found the peer review process quick and positive in its input. The support from the editorial officer has been very agile, always with the intention of improving the article and taking into account our subsequent corrections.

My article, titled 'No Way Out of the Smartphone Epidemic Without Considering the Insights of Brain Research,' has been republished in the International Journal of Clinical Case Reports and Reviews. The review process was seamless and professional, with the editors being both friendly and supportive. I am deeply grateful for their efforts.
To Dear Erin Aust – Editorial Coordinator of Journal of General Medicine and Clinical Practice! I declare that I am absolutely satisfied with your work carried out with great competence in following the manuscript during the various stages from its receipt, during the revision process to the final acceptance for publication. Thank Prof. Elvira Farina

Dear Jessica, and the super professional team of the ‘Clinical Cardiology and Cardiovascular Interventions’ I am sincerely grateful to the coordinated work of the journal team for the no problem with the submission of my manuscript: “Cardiometabolic Disorders in A Pregnant Woman with Severe Preeclampsia on the Background of Morbid Obesity (Case Report).” The review process by 5 experts was fast, and the comments were professional, which made it more specific and academic, and the process of publication and presentation of the article was excellent. I recommend that my colleagues publish articles in this journal, and I am interested in further scientific cooperation. Sincerely and best wishes, Dr. Oleg Golyanovskiy.

Dear Ashley Rosa, Editorial Coordinator of the journal - Psychology and Mental Health Care. " The process of obtaining publication of my article in the Psychology and Mental Health Journal was positive in all areas. The peer review process resulted in a number of valuable comments, the editorial process was collaborative and timely, and the quality of this journal has been quickly noticed, resulting in alternative journals contacting me to publish with them." Warm regards, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. I appreciate the journal (JCCI) editorial office support, the entire team leads were always ready to help, not only on technical front but also on thorough process. Also, I should thank dear reviewers’ attention to detail and creative approach to teach me and bring new insights by their comments. Surely, more discussions and introduction of other hemodynamic devices would provide better prevention and management of shock states. Your efforts and dedication in presenting educational materials in this journal are commendable. Best wishes from, Farahnaz Fallahian.
Dear Maria Emerson, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. I am delighted to have published our manuscript, "Acute Colonic Pseudo-Obstruction (ACPO): A rare but serious complication following caesarean section." I want to thank the editorial team, especially Maria Emerson, for their prompt review of the manuscript, quick responses to queries, and overall support. Yours sincerely Dr. Victor Olagundoye.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. Many thanks for publishing this manuscript after I lost confidence the editors were most helpful, more than other journals Best wishes from, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Agrippa Hilda, Editorial Coordinator, Journal of Neuroscience and Neurological Surgery. The entire process including article submission, review, revision, and publication was extremely easy. The journal editor was prompt and helpful, and the reviewers contributed to the quality of the paper. Thank you so much! Eric Nussbaum, MD
Dr Hala Al Shaikh This is to acknowledge that the peer review process for the article ’ A Novel Gnrh1 Gene Mutation in Four Omani Male Siblings, Presentation and Management ’ sent to the International Journal of Clinical Case Reports and Reviews was quick and smooth. The editorial office was prompt with easy communication.

Dear Erin Aust, Editorial Coordinator, Journal of General Medicine and Clinical Practice. We are pleased to share our experience with the “Journal of General Medicine and Clinical Practice”, following the successful publication of our article. The peer review process was thorough and constructive, helping to improve the clarity and quality of the manuscript. We are especially thankful to Ms. Erin Aust, the Editorial Coordinator, for her prompt communication and continuous support throughout the process. Her professionalism ensured a smooth and efficient publication experience. The journal upholds high editorial standards, and we highly recommend it to fellow researchers seeking a credible platform for their work. Best wishes By, Dr. Rakhi Mishra.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. The peer review process of the journal of Clinical Cardiology and Cardiovascular Interventions was excellent and fast, as was the support of the editorial office and the quality of the journal. Kind regards Walter F. Riesen Prof. Dr. Dr. h.c. Walter F. Riesen.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. Thank you for publishing our article, Exploring Clozapine's Efficacy in Managing Aggression: A Multiple Single-Case Study in Forensic Psychiatry in the international journal of clinical case reports and reviews. We found the peer review process very professional and efficient. The comments were constructive, and the whole process was efficient. On behalf of the co-authors, I would like to thank you for publishing this article. With regards, Dr. Jelle R. Lettinga.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, I would like to express my deep admiration for the exceptional professionalism demonstrated by your journal. I am thoroughly impressed by the speed of the editorial process, the substantive and insightful reviews, and the meticulous preparation of the manuscript for publication. Additionally, I greatly appreciate the courteous and immediate responses from your editorial office to all my inquiries. Best Regards, Dariusz Ziora

Dear Chrystine Mejia, Editorial Coordinator, Journal of Neurodegeneration and Neurorehabilitation, Auctores Publishing LLC, We would like to thank the editorial team for the smooth and high-quality communication leading up to the publication of our article in the Journal of Neurodegeneration and Neurorehabilitation. The reviewers have extensive knowledge in the field, and their relevant questions helped to add value to our publication. Kind regards, Dr. Ravi Shrivastava.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, Auctores Publishing LLC, USA Office: +1-(302)-520-2644. I would like to express my sincere appreciation for the efficient and professional handling of my case report by the ‘Journal of Clinical Case Reports and Studies’. The peer review process was not only fast but also highly constructive—the reviewers’ comments were clear, relevant, and greatly helped me improve the quality and clarity of my manuscript. I also received excellent support from the editorial office throughout the process. Communication was smooth and timely, and I felt well guided at every stage, from submission to publication. The overall quality and rigor of the journal are truly commendable. I am pleased to have published my work with Journal of Clinical Case Reports and Studies, and I look forward to future opportunities for collaboration. Sincerely, Aline Tollet, UCLouvain.

Dear Ms. Mayra Duenas, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. “The International Journal of Clinical Case Reports and Reviews represented the “ideal house” to share with the research community a first experience with the use of the Simeox device for speech rehabilitation. High scientific reputation and attractive website communication were first determinants for the selection of this Journal, and the following submission process exceeded expectations: fast but highly professional peer review, great support by the editorial office, elegant graphic layout. Exactly what a dynamic research team - also composed by allied professionals - needs!" From, Chiara Beccaluva, PT - Italy.

Dear Maria Emerson, Editorial Coordinator, we have deeply appreciated the professionalism demonstrated by the International Journal of Clinical Case Reports and Reviews. The reviewers have extensive knowledge of our field and have been very efficient and fast in supporting the process. I am really looking forward to further collaboration. Thanks. Best regards, Dr. Claudio Ligresti
Dear Chrystine Mejia, Editorial Coordinator, Journal of Neurodegeneration and Neurorehabilitation. “The peer review process was efficient and constructive, and the editorial office provided excellent communication and support throughout. The journal ensures scientific rigor and high editorial standards, while also offering a smooth and timely publication process. We sincerely appreciate the work of the editorial team in facilitating the dissemination of innovative approaches such as the Bonori Method.” Best regards, Dr. Matteo Bonori.
